Ronald Gangnon
Professor of Biostatistics
University of Wisconsin-Madison
Biography
I am a Professor in the Department of Biostatistics and Medical Informatics and the Department of Population Health Sciences in the School of Medicine and Public Health at the University of Wisconsin-Madison. I have an affiliate appointment in the Department of Statistics.
I grew up in Duluth, Minnesota. I graduated from East High School in 1988. I received a BA in Mathematics and Economics in 1992 from the University of Minnesota-Duluth and an MS in Statistics in 1994 and a PhD in Statistics with emphasis in Biostatistics in 1998 from the University of Wisconsin-Madison. My PhD advisor was Murray Clayton.
I was a research scientist in the Statistical Data Analysis Center (SDAC) in the Department of Biostatistics and Medical Informatics at the University of Wisconsin-Madison, 1998-2005. I joined the faculty with a joint appointment in the Department of Biostatistics and Medical Informatics and the Department of Population Health Sciences in 2005.
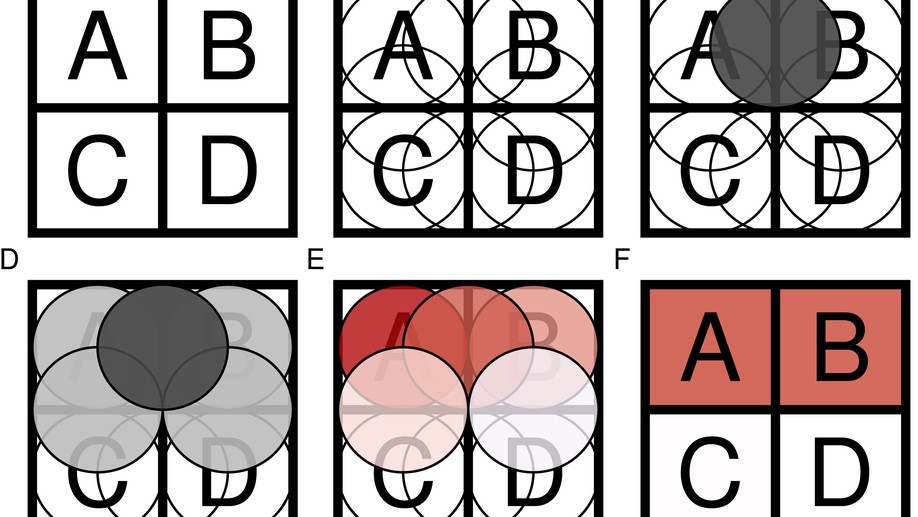
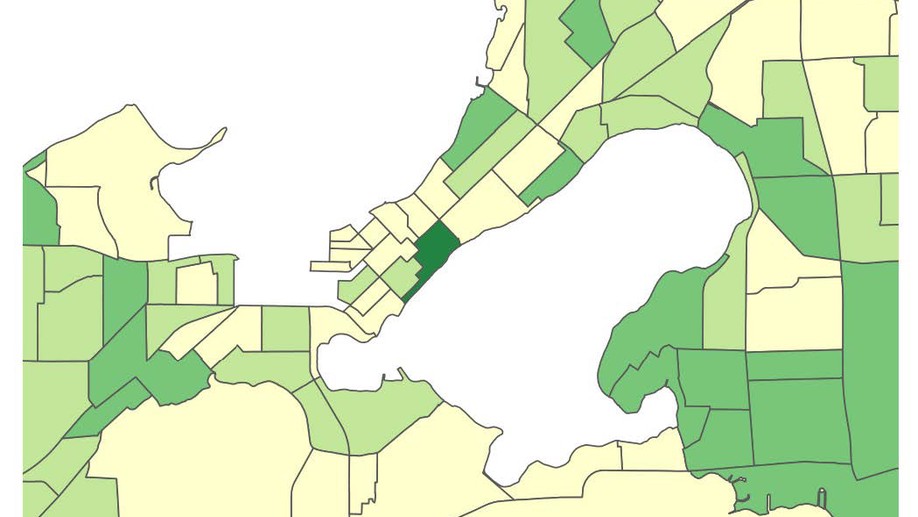
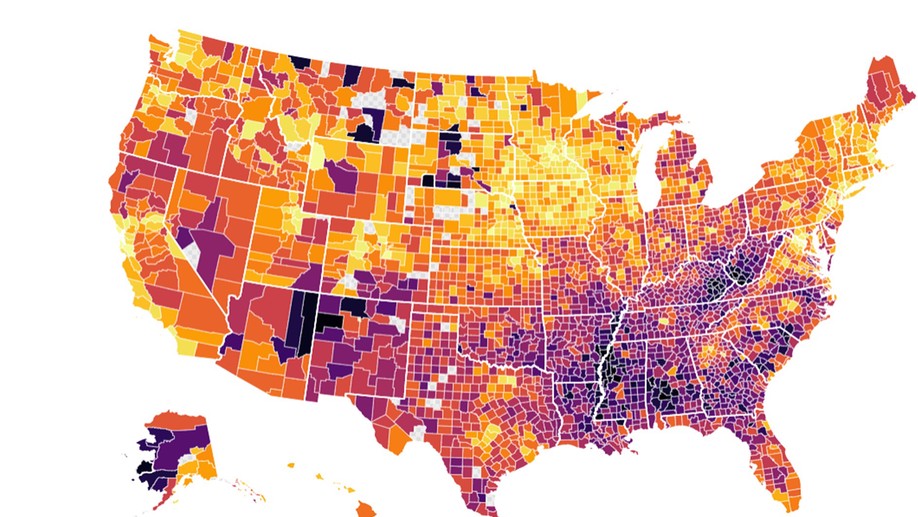
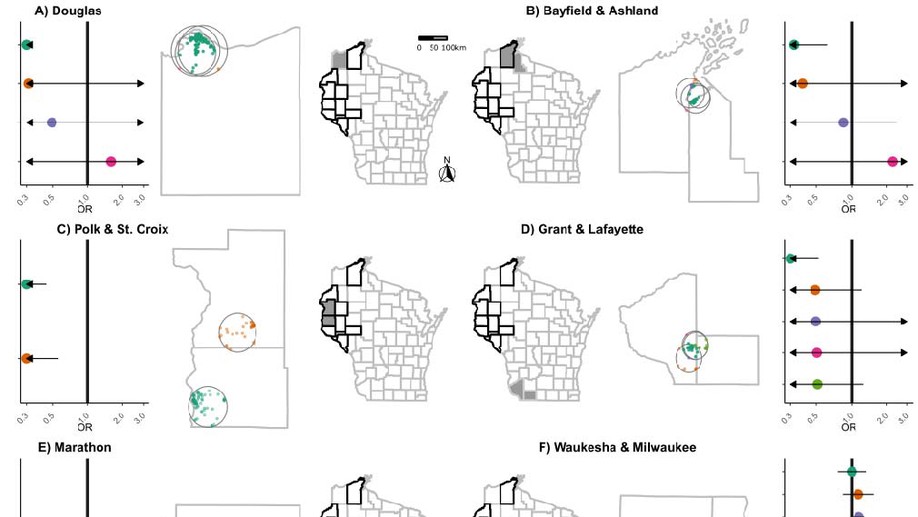
I am an applied biostatistician focusing on problems in clinical and epidemiologic research. Current methodologic areas of interest include (1) multi-state models for incidence, progression and regression of ocular (and other) diseases, (2) small area estimation problems, particularly ranking, (3) spatial and spatio-temporal modeling, particularly cluster detection and high-dimensional variable selection and (4) age-period-cohort modeling.
Outside of the office, I’m an avid cyclist. You can check out my recent rides on Strava. I’m also a big movie fan. You’ll definitely see me at the Wisconsin Film Festival, one of my favorite events every year, and you can see lists of my favorite films and what I’ve been watching recently on Letterboxd.
Interests
- Spatial and Spatio-Temporal Modeling
- Age-Period-Cohort Models
- Ranking
- Multi-State Models
Education
-
PhD in Statistics (emphasis in Biostatistics), 1998
University of Wisconsin-Madison
-
MS in Statistics, 1994
University of Wisconsin-Madison
-
BA in Mathematics and Economics, 1992
University of Minnesota-Duluth